- How to run an existing protocol?
- How does it works?
- A Hello World! example.
- How to modify an existing protocol?
- Now to write a new protocol?
How to run an existing protocol?¶
Make sure you have a working python3 interprete in your device (PC, tablet, smartphone, etc.) and a copy of RobotEvo (downloaded or cloned from GitHub).
Now, the simplest way is to run the python script containing the protocol, providing it have a “main” function.
For better control, in a your onw python script, import the desired protocol and just create an instance (python object) of the protocol, possibly setting some of the constructor parameters, and call the .run() method of that object. You can see many examples of this usage in the script robotevo/protocols/test.py This will create a set of files with the generated evoware scripts, a human readable protocol, and comments, possibly including warnings. It may abort with more or less detailed messages about the errors.
Alternatively run GUI.py (only in devices with a functioning standard python module “tkinter”) to select the protocol, the desired protocol “variant” and change some other minor parameters like number of samples or required reagents volume.
To make the actual pippeting in the real robot, open the generated .esc script in the EVOware editor. It will alert you that the check sum have not been set, which in this case just flags the fact that this is a newly generated script you have not run yet. Accept to load it in the EVOware script editor. Here you will have very good assistance to visualize the details of each step and to do a normal, full TECAN validation of the correctness of the script.
Use the information from the visual worktable map to physically setup the labware. Use the detailed comments automatically inserted by RobotEvo in the script or the associated .protocol.txt file to fill the expected initial volume of each reagent.
Use EVOware to run the script as usually.
How does it works?¶
Already the creation of the protocol object will run some “boilerplate” code to setup things we need to run the useful part of our protocol.
For example it will parse the provided worktable template file (a .ewt or just a compatible .esc evoware file) and will remember (in a sort of map) all the labware present in the worktable, including its unique name, type and location.
It will also initialize some other characteristics of the used robot (not present in the worktable file) like number of tips in the LiHa, etc.
Additionally it will set the desired EvoMode: what kind of output we want to produce - normally an evoware script (his generated script will include again all the information for the worktable), but also a human readable protocol, etc.
By running .run() we “create” or “get”, from the parsed worktable file, labwares, like multiplates, tube racks, etc, and “create” the reagents defined there in the script, including location in the worktable, volume, etc. This make possible for the “internal iRobot” to model or track the content of each well, and to detect (and report) potential logical errors in the protocol.
If at this point the protocol include a call to .cehcklist() instructions will be generated to inform to the human robot-operator at run time, the positions and initial quantity of all reagents he need to make sure are in place. If a GUI is in use and was previosly created a new sub-GUI will be automatically generated to show all the details of the defined reagents making possible to change some properties without modifying programmatically the protocol.
A typical protocol will use the high level instructions inherited from protocol_steps, like transfer, distribute, with tips, etc., to express the “physical” protocol. This instructions are in turn internally implemented using lower level instructions like aspirate, get tips. etc. Each of this low level intructions will interact with the selected EvoMode to generate the corresponding instructions in the EVOware script and to check errors and change the state of the internally modeled iRobot, including the liquid volume in each well and tip and many other details.
A Hello World! example.¶
Let create the classical, in the the world of programming, Hello World! example. It will just shows that message in the screen of the PC controlling the robot and will wait for user confirmation producing a typical sound.
By running the script:
#
"""
Hello World!
------------
The classical, in the the world of programming, Hello World! example.
It will just shows that message in the screen of the PC controlling the robot and will wait
for user confirmation producing a typical sound.
"""
from EvoScriPy.protocol_steps import *
class HelloWorld(Protocol):
"""
This is a very general protocol.
Normally you will inherit from a Protocol class adapted to your real robot.
"""
name = "Hello World"
def __init__(self, GUI = None,
output_filename = None,
worktable_template_filename = None):
this = Path(__file__).parent
Protocol.__init__(self,
GUI = GUI,
output_filename = output_filename or this / 'scripts' / 'hello_world',
worktable_template_filename = worktable_template_filename or this / 'hello_world.ewt')
def run(self):
self.check_list()
self.user_prompt("Hello World!")
self.done()
if __name__ == "__main__":
HelloWorld().run()
[IMPORTANT: replace the worktable_template_filename argument with any valid -for your very onw robot- worktable template (.ewt) or script (.esc).]
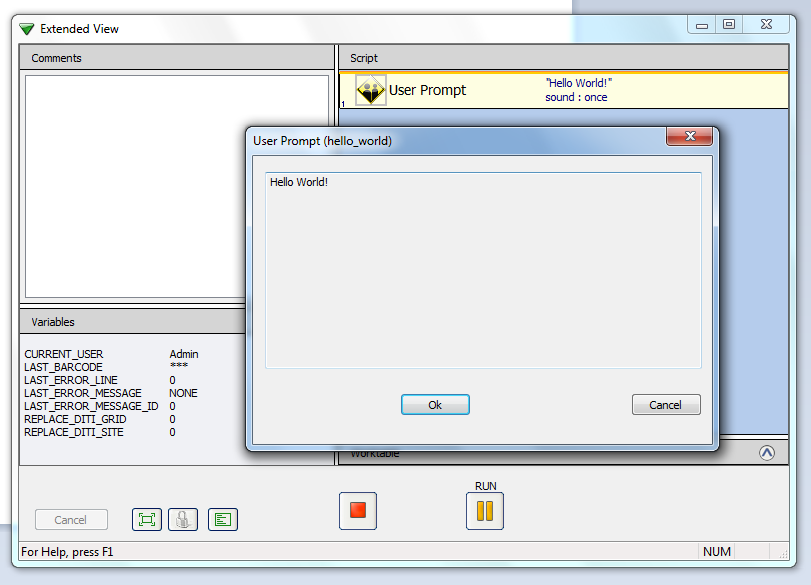
we will have some files (currently 4) generated with names following the pattern of the output_filename constructor argument: in particular ‘../current/tests/hello_world.esc’ will contain a new evoware script you can load into the Freedom evoware editor. After you agree to use the script with an still unvalidated check-summe you will see it just contain an instruction for a simple user promt. By using evoware to run this script you will get:
How to modify an existing protocol?¶
How to write a new protocol?¶
# author Ariel Vina-Rodriguez (qPCR4vir)
# this example is free to use at your onw risk
# 2019-2019
"""
Prepare serial dilutions of two mixes.
--------------------------------------
"""
__author__ = 'Ariel'
# Tutorial
from protocols.evo200_f.evo200_f import *
class DemoTwoMixes(Evo200_FLI):
"""
Prepare two 1:10 serial dilutions of two different mixes each in 'n' 100 uL final volume wells
(each in a microplate, the second one to be moved to the working position).
'mix1' and 'mix2' are diluted separately in n wells 1:10 (mix1_10 and mix2_10 respectively) using
a provided "buffer". From those wells a portion is transferred to the final 1:100 dilutions
(mix1_100 and mix2_100 respectively) to fv=100 uL final volume
One way to achieve this:
- Calculate how much to transfer from each mix1_10 to mix1_100. v_mix1_10_100= fv/10 and from diluent.
- Calculate how much to distribute from mix1 to each mix1_10 and from diluent.
- Define a reagent `mix1` and `mix2`in an Eppendorf rack (labware) for the calculated volume per "sample" (mix1_10 or 2).
- Define a reagent `buffer` in a 100 mL cubette `BufferCub` for the total volume per "sample".
- Generate check list
- Transfer plate 2 from the original location `plate2` to the final location `plate2-moved`
- Define derived reagents for diluted mixes
- Distribute mix1 and buffer into mix1_10 and similar with mix2
- Transfer from mix1_10 to mix1_100 and distribute buffer here. The same with mix2_10
"""
name = "Prefill one plate with Buffer."
min_s, max_s = 1, 96/2
# for now just ignore the variants
def def_versions(self):
self.versions = {'No version': self.V_def}
def V_def(self):
pass
def __init__(self,
GUI = None,
num_of_samples: int = None,
worktable_template_filename = None,
output_filename = None,
first_tip = None,
run_name: str = ""):
this = Path(__file__).parent
Evo200_FLI.__init__(self,
GUI = GUI,
num_of_samples = num_of_samples or DemoTwoMixes.max_s,
worktable_template_filename = worktable_template_filename or
this / 'demo-two.mixes.Evo200example.ewt',
output_filename = output_filename or this / 'scripts' / 'two.mixes',
first_tip = first_tip,
run_name = run_name)
def run(self):
self.initialize() # set_EvoMode and set_defaults() from Evo200
self.check_initial_liquid_level = True
self.show_runtime_check_list = True
num_of_samples = self.num_of_samples
assert self.min_s <= num_of_samples <= self.max_s, "In this demo we want to set 2x num_of_samples in a 96 well plate."
wt = self.worktable
self.comment('Prefill a plate with some dilutions of two master mix and Buffer Reagent for {:d} samples.'
.format(num_of_samples))
buf_cuvette = wt.get_labware("BufferCub", labware.Trough_100ml) # Get Labwares from the work table
master_mixes_ = wt.get_labware("mixes", labware.Eppendorfrack)
buf_per_sample =0
fv = 100
v_mix_10_100 = fv / 10 # to be transferred from mix1_10 to mix1_100
buf_mix_100 = fv - v_mix_10_100
buf_per_sample += buf_mix_100
v_mix_10 = (fv + v_mix_10_100)/10 # to be distribute from original mix1 to mix1_10
buf_mix_10 = (fv + v_mix_10_100) - v_mix_10
buf_per_sample += buf_mix_10
# Define the reagents in each labware (Cuvette, eppys, etc.)
buffer = Reagent("Buffer ", buf_cuvette, volpersample = buf_per_sample,
def_liq_class = self.Water_wet,
num_of_samples = 2 * self.num_of_samples)
mix1 = Reagent("mix1", master_mixes_, volpersample = v_mix_10,
def_liq_class = self.Water_wet,
num_of_samples = self.num_of_samples)
mix2 = Reagent("mix2", master_mixes_, volpersample = v_mix_10,
def_liq_class = self.Water_wet,
num_of_samples = self.num_of_samples)
# Show the check_list GUI to the user for possible small changes
self.check_list()
instructions.wash_tips(wasteVol=5, FastWash=True).exec()
plate1 = wt.get_labware("plate1", '96 Well Microplate')
plate2 = wt.get_labware("plate2", '96 Well Microplate')
new_location = wt.get_labware("plate2-moved").location
Reagent.use_minimal_number_of_aliquots = False # Define derived reagents ---------------------
mix1_10 = Reagent(f"mix1, diluted 1:10",
plate1,
initial_vol = 0.0,
num_of_aliquots= num_of_samples,
def_liq_class = self.Water_free,
excess = 0)
mix2_10 = Reagent(f"mix2, diluted 1:10",
plate2,
initial_vol = 0.0,
num_of_aliquots= num_of_samples,
def_liq_class = self.Water_free,
excess = 0)
mix1_100 = Reagent(f"mix1, diluted 1:100",
plate1,
wells = 'A07',
initial_vol = 0.0,
num_of_aliquots= num_of_samples,
def_liq_class = self.Water_free,
excess = 0)
mix2_100 = Reagent(f"mix2, diluted 1:100",
plate2,
wells = 'A07',
initial_vol = 0.0,
num_of_aliquots= num_of_samples,
def_liq_class = self.Water_free,
excess = 0)
instructions.transfer_rack(plate2, new_location).exec() # just showing how RoMa works.
with group("Fill plate with mixes "):
self.user_prompt("Put the plates for Buffer ")
with self.tips(reuse=True, drop=False):
self.distribute(reagent = mix1,
to_labware_region = mix1_10.select_all())
with self.tips(reuse=True, drop=False):
self.distribute(reagent = mix2,
to_labware_region = mix2_10.select_all())
with self.tips(reuse=True, drop=False):
self.distribute(reagent=buffer, to_labware_region=mix1_10.select_all(), volume=buf_mix_10)
self.distribute(reagent=buffer, to_labware_region=mix2_10.select_all(), volume=buf_mix_10)
with self.tips(reuse=True, drop=False):
wells_100 = mix1_100.select_all()
self.transfer(from_labware_region = mix1_10.select_all(),
to_labware_region = wells_100,
volume = v_mix_10_100)
with self.tips(reuse=True, drop=False):
wells_100 = mix2_100.select_all()
self.transfer(from_labware_region = mix2_10.select_all(),
to_labware_region = wells_100,
volume = v_mix_10_100)
with self.tips(reuse=True, drop=False):
self.distribute(reagent=buffer, to_labware_region=mix1_100.select_all(), volume=buf_mix_100)
self.distribute(reagent=buffer, to_labware_region=mix2_100.select_all(), volume=buf_mix_100)
self.drop_tips()
self.done()
if __name__ == "__main__":
p = DemoTwoMixes(num_of_samples= 4,
run_name= "_4s_mix_1_2")
p.use_version('No version')
# p.set_first_tip('A01')
p.run()
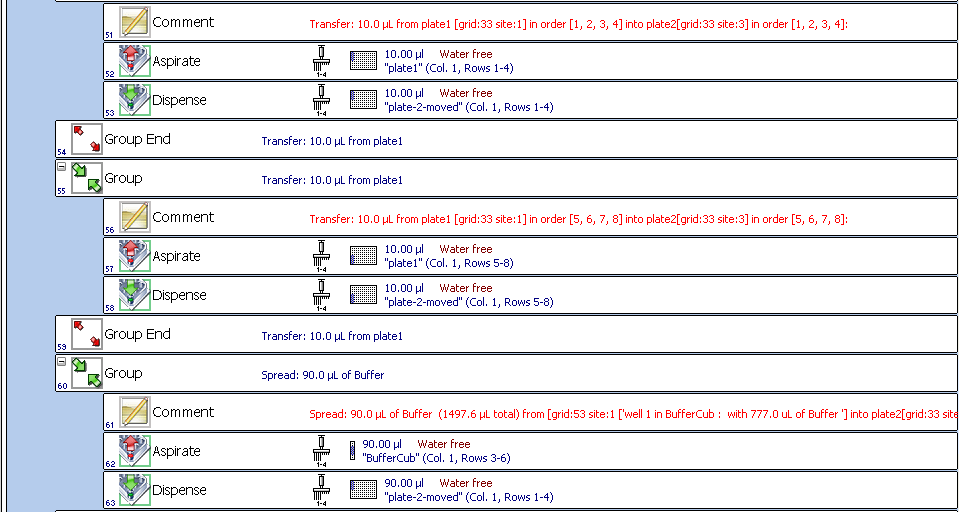
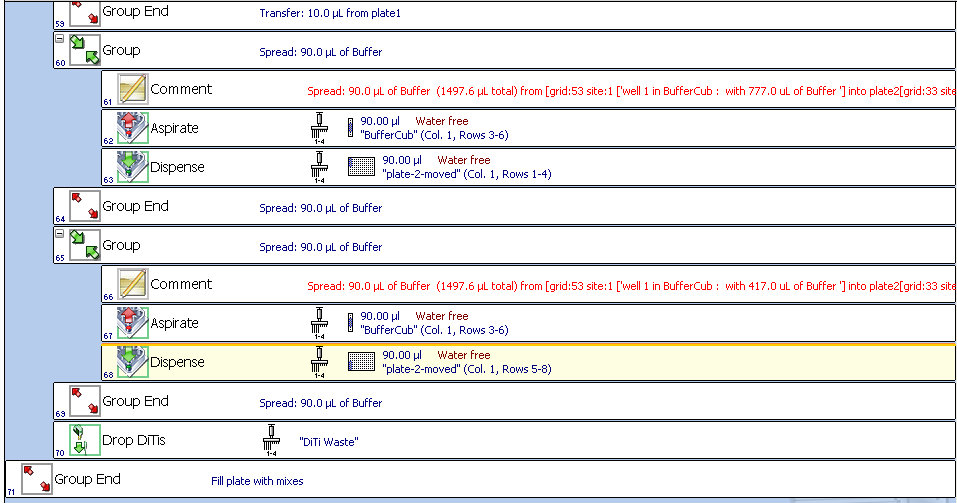
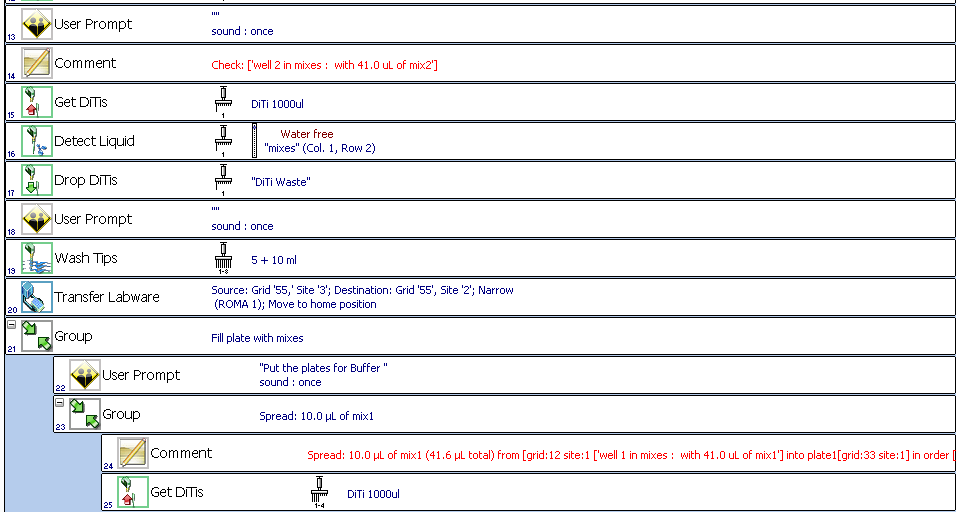
we will have something like: